GSEA v2.08. Release Notes
<a href="http://www.broadinstitute.org/gsea/">GSEA Home</a> | <a href="http://www.broadinstitute.org/gsea/downloads.jsp">Downloads</a> | <a href="http://www.broadinstitute.org/gsea/msigdb/">Molecular Signatures Database</a> | Documentation | <a href="http://www.broadinstitute.org/gsea/contact.jsp">Contact</a>
In release 2.0.8, there is no change to the GSEA algorithm.
The changes include:
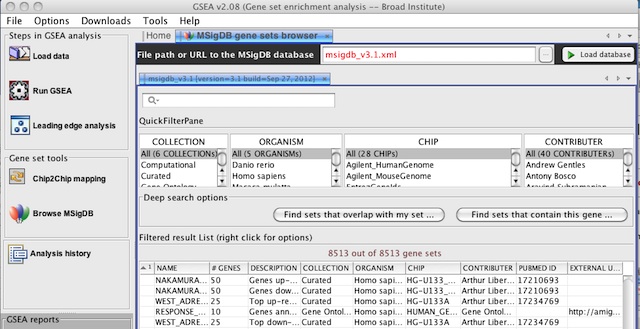
- When you Browse MSigDB, by default it will now display version 3.1 of the MSigDB gene sets.
To browse earlier versions of MSigDB, type msigdb_v3.xml or msigdb_v2.5.xml in the File path or URL to the MSigDB database and click the Load database button.
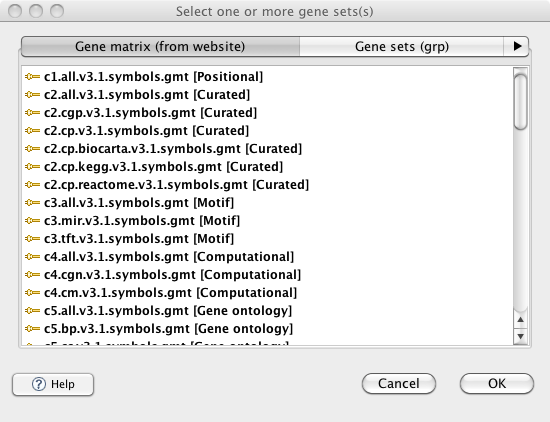
In the previous version of GSEA, there were display issues when viewing the selection menu of GMT files in the MSigDB database from the Run Gsea screen and the Chip2Chip mapping screen. These issues have been fixed. The GMT files for the latest release of MSigDB are now always displayed at the top of the list and in a bold font.
The latest version of MSigDB is significantly larger than previous versions and we now recommend that you run GSEA with no less than 1 GB of memory. If you run GSEA from the command line, use the -Xmx Java option to specify increased heap size. For example, java -Xmx1024m -jar gsea2-2.08.jar.