SNP2HLA: Imputation of Amino Acid Polymorphisms in Human Leukocyte Antigens
SNP2HLA is a tool to impute amino acid polymorphisms and single nucleotide polymorphisms in human luekocyte antigenes (HLA) within the major histocompatibility complex (MHC) region in chromosome 6.
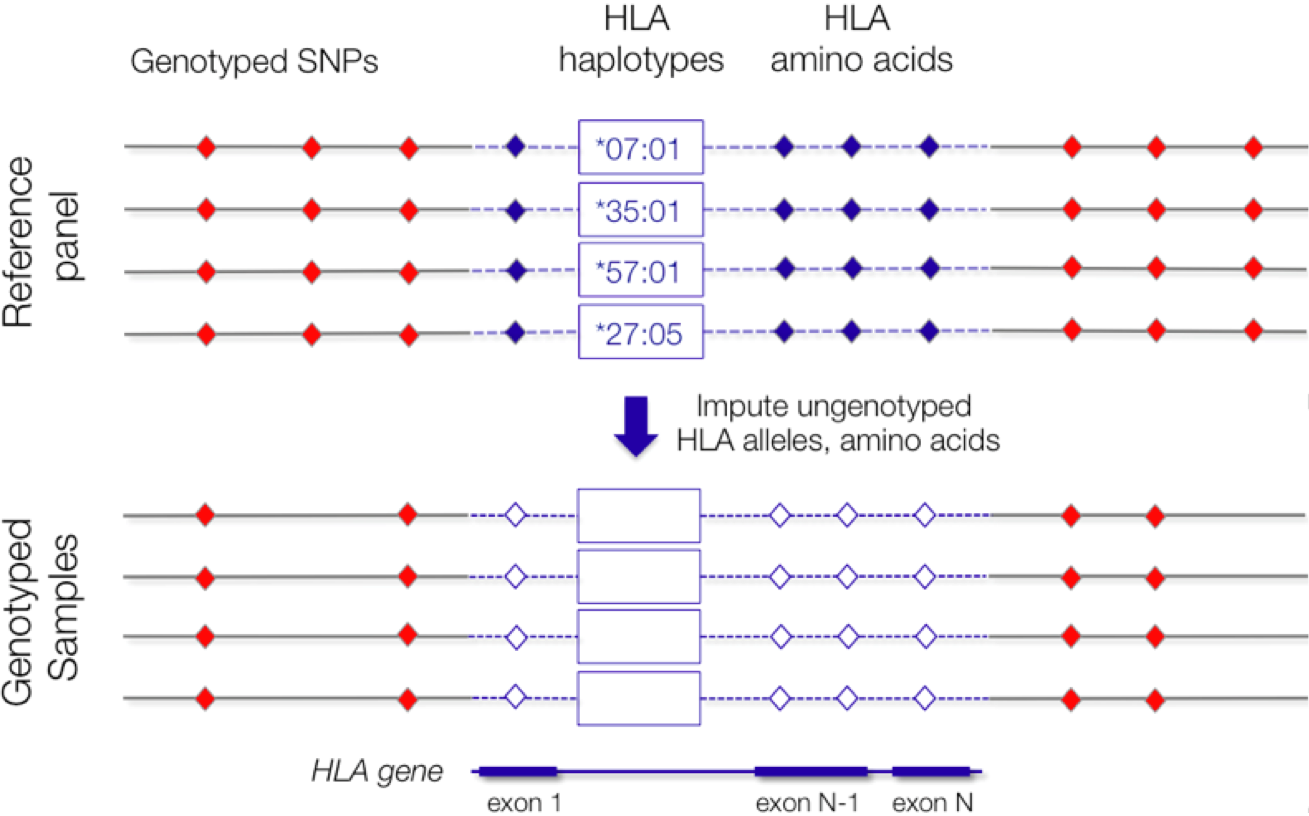
The unique feature of SNP2HLA is that it imputes not only the classical HLA alleles but also the amino acid sequences of those classical alleles, so that individual amino acid sites can be directly tested for association. This allows for facile amino-acid focused downstream analysis.
SNP2HLA also provides a companion package, MakeReference. This is software that builds the reference panel that can be used for SNP2HLA. This package is used for the situation that the provided reference panel is inappropriate (e.g. different populations), and the user wants to build the reference panel by him(her)self, e.g. typing the HLA alleles in a subset of individuals.
SNP2HLA is developed by Sherman Jia and Buhm Han in the labs of Soumya Raychaudhuri and Paul de Bakker at the Brigham and Women's Hospital and Harvard Medical School, and the Broad Institute.
Links
-
Download: SNP2HLA v1.0.3 package including SNP2HLA, MakeReference, and Pan Asian Reference Panel (tarball)
----- NEWS1: Pan-Asian reference panel is now included in the package V1.0.2! (7/10/14)
----- NEWS2: T1DGC reference panel is now removed from the package V1.0.3 due to security issues relating to individual-level genotype data (3/10/15). If you are a researcher interested in obtaining access to this reference panel, please contact the NIDDK Central Repository ( https://repository.niddk.nih.gov/studies/t1dgc-special/?query=snp2hla)
- Manual: SNP2HLA
- Manual: MakeReference
Reference
If you use the Pan-Asian reference panel, please cite the following papers:
1. Pillai NE et al. "Predicting HLA alleles from high-resolution SNP data in three Southeast Asian populations." Hum Mol Genet, [epub ahead of print] doi:10.1093/hmg/ddu149. 2014.
2. Okada Y et al. (2014) Risk for ACPA-positive rheumatoid arthritis is driven by shared HLA amino acid polymorphisms in Asian and European populations. Hum Mol Genet, In press.
Any questions?
You can contact us at snp2hla AT broadinstitute DOT org