BUHMBOX: Distinguishing pleiotropy and heterogeneity
BUHMBOX is a tool to distinguish pleiotropy and heterogeneitiy. Suppose that we collect disease A cases. Then we observe that known disease B risk alleles are enriched within disease A cases. Is this because disease A and B have pleiotropy, or because disease A patients have a subset that shows genetic characteristics of disease B? BUHMBOX (Breaking Up Heterogeneous Mixture Based On Cross-locus correlations) is a tool to distinguish these two situations.
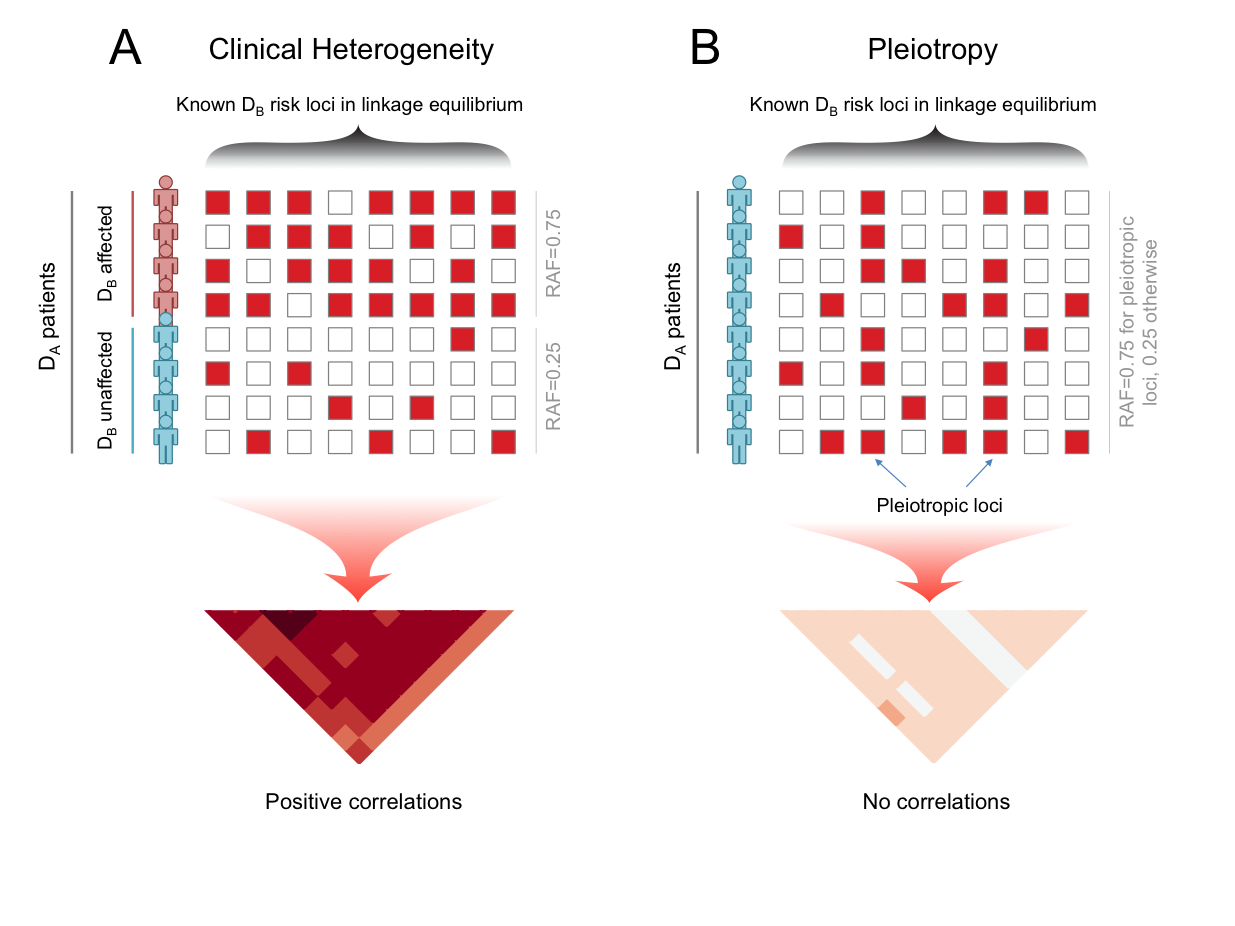
The intuition under BUHMBOX is that we will observe weak positive correlations between disease-B-loci in disease A cases, if it were due to heterogeneity, because of the subgrouping. Thus BUHMBOX aggregrates the correlations into one statistic. To strictly control for false positives, BUHMBOX uses pruning, PCs, and comparison to controls.
BUHMBOX is developed by Buhm Han in the labs of Soumya Raychaudhuri and Paul de Bakker at the Brigham and Women's Hospital and Harvard Medical School, and the Broad Institute.
Links
- Download: BUHMBOX v0.38 (R script) <--- right click and download
- Download: BUHMBOX power calculation code (R script) <--- right click and download
- Manual: BUHMBOX
- Manual: BUHMBOX Power calculation
Reference
Also was at Biorxiv: http://biorxiv.org/content/early/2015/11/06/030783
Any questions?
You can contact us at buhmhan AT amc DOT seoul DOT kr